Science Highlights
Using Machine Learning to Identify a Signature of Bipolar Disorder
 In collaboration with researchers from the Big Data to Knowledge (BD2K) ENIGMA Center, Abraham Nunes MD., M.B.A., and Tomas Hajek M.D., Ph.D., published a study evaluating machine-learning methods to identify diagnostic markers of bipolar disorder. "After assembling over 3,000 brain scans of bipolar disorder patients and controls from multiple sites, the researchers tested whether computers could train themselves to reliably distinguish patients from controls. The results, considered promising, provide a proof-of-concept for a brain-imaging signature of bipolar disorder to identify the illness in individuals," noted a news feature on the initiative, by the Brain & Behavior Research Foundation. The study, recently published in Molecular Psychiatry, reports the latest results from an international alliance to understand what brain features might consistently distinguish bipolar patients from healthy individuals.
In collaboration with researchers from the Big Data to Knowledge (BD2K) ENIGMA Center, Abraham Nunes MD., M.B.A., and Tomas Hajek M.D., Ph.D., published a study evaluating machine-learning methods to identify diagnostic markers of bipolar disorder. "After assembling over 3,000 brain scans of bipolar disorder patients and controls from multiple sites, the researchers tested whether computers could train themselves to reliably distinguish patients from controls. The results, considered promising, provide a proof-of-concept for a brain-imaging signature of bipolar disorder to identify the illness in individuals," noted a news feature on the initiative, by the Brain & Behavior Research Foundation. The study, recently published in Molecular Psychiatry, reports the latest results from an international alliance to understand what brain features might consistently distinguish bipolar patients from healthy individuals.
Read the news coverage from the Brain & Behavior Research Foundation
Reference:
Using structural MRI to identify bipolar disorders - 13 site machine learning study in 3020 individuals from the ENIGMA Bipolar Disorders Working Group. Abraham Nunes, Tomas Hajek, et al. Mol Psychiatry. 2018 Aug 31. doi: 10.1038/s41380-018-0228-9
Using LINCS and other Big Data to Find New Cancer Treatments
 A new study using computational tools to combine multiple sets of previously generated biomedical big data has identified drugs that can be repurposed to fight cancer. Dr. Bin Chen, a Big Data to Knowledge (BD2K) Mentored Career Development Awardee, and colleagues took advantage of multiple existing data sets, including the NIH Common Fund’s Library of Integrated Network-based Cellular Signatures (LINCS) L1000 data set and the National Cancer Institute’s The Cancer Genome Atlas (TCGA) data set to generate a list of potential drugs predicted to inhibit the growth of liver cancer cells. Dr. Chen and colleagues compared the global cellular changes between normal and cancerous liver cells in the TCGA data set with changes caused by treating cells with a panel of 12,442 distinct chemical compounds in the LINCS data sets. They tested the hypothesis that compounds causing inverse changes to those seen when a cell becomes cancerous would be strong candidates for treating cancer. Using this approach, four compounds were selected that had the potential to reverse the changes seen in liver cancer patients. When these compounds were tested against liver cancer cell lines grown in the laboratory, all four inhibited the growth of these cells. The most potent of these drugs was also tested in a mouse liver cancer model and was shown to reduce the size of tumors. This drug (pyrvinium pamoate), which has been previously FDA approved to treat pinworm infections, will require further experimentation to determine if it can be repurposed to treat liver cancer in humans.
A new study using computational tools to combine multiple sets of previously generated biomedical big data has identified drugs that can be repurposed to fight cancer. Dr. Bin Chen, a Big Data to Knowledge (BD2K) Mentored Career Development Awardee, and colleagues took advantage of multiple existing data sets, including the NIH Common Fund’s Library of Integrated Network-based Cellular Signatures (LINCS) L1000 data set and the National Cancer Institute’s The Cancer Genome Atlas (TCGA) data set to generate a list of potential drugs predicted to inhibit the growth of liver cancer cells. Dr. Chen and colleagues compared the global cellular changes between normal and cancerous liver cells in the TCGA data set with changes caused by treating cells with a panel of 12,442 distinct chemical compounds in the LINCS data sets. They tested the hypothesis that compounds causing inverse changes to those seen when a cell becomes cancerous would be strong candidates for treating cancer. Using this approach, four compounds were selected that had the potential to reverse the changes seen in liver cancer patients. When these compounds were tested against liver cancer cell lines grown in the laboratory, all four inhibited the growth of these cells. The most potent of these drugs was also tested in a mouse liver cancer model and was shown to reduce the size of tumors. This drug (pyrvinium pamoate), which has been previously FDA approved to treat pinworm infections, will require further experimentation to determine if it can be repurposed to treat liver cancer in humans.
The growing number of large, robust experimental data sets in biomedical research offers unprecedented opportunities for new discoveries using computational approaches, which is a continuing goal of the BD2K program. This study, also demonstrates the power of using the LINCS data sets to discover novel therapeutics by focusing on global cellular changes instead of a singular molecular target. Similar approaches have the potential to yield candidate drugs for repurposing to treat a wide spectrum of diseases.
Read the press release from UCSF here.
Reference:
Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets. Chen B, Ma L, Paik H1, Sirota M, Wei W, Chua MS, So S, Butte AJ. Nat Commun. 2017 Jul 12. 8:16022.
The ENIGMA Consortium Maps the Brain’s Gray Matter in People with Bipolar Disorder
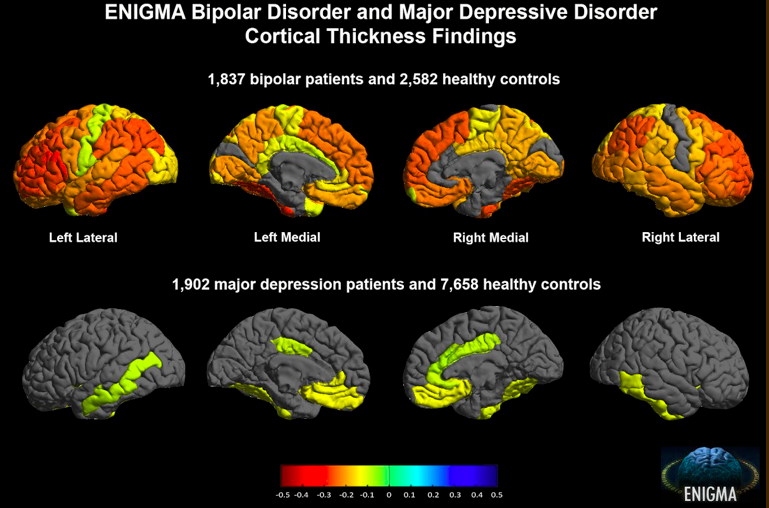
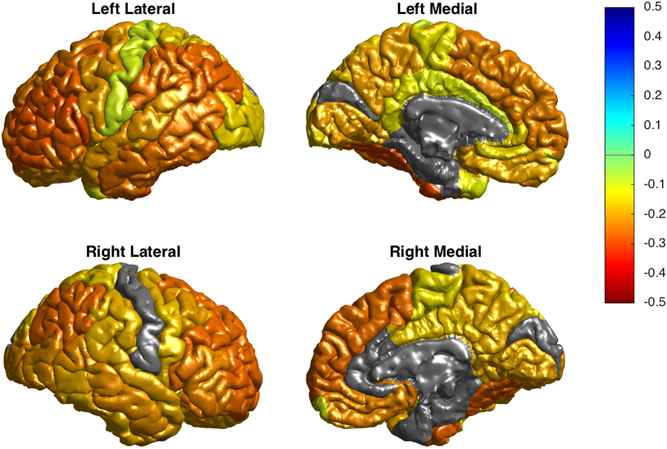 Bipolar disorder, which affects 1-3% of the adult population, is a disorder of the brain that results in extreme shifts in mood from periods being extremely “up” (manic episodes) to extremely “down” (depressive episodes). Despite bipolar disorder being a common disease, not much is known about the biological mechanisms that lead to it.
Bipolar disorder, which affects 1-3% of the adult population, is a disorder of the brain that results in extreme shifts in mood from periods being extremely “up” (manic episodes) to extremely “down” (depressive episodes). Despite bipolar disorder being a common disease, not much is known about the biological mechanisms that lead to it.
In the largest ever imaging study of bipolar disorder, the ENIGMA consortium, a BD2K Center of Excellence, found that the gray matter in particular sections of the brain in people with bipolar disorder is thinner than in healthy individuals. Gray matter covers the surface of the brain and is where the bodies of the brain’s nerve cells are located. This study, which was a collaboration between 79 institutions worldwide, analyzed the MRI scans of 6503 individuals in order to control for variables such as age, sex, prescribed medications, and duration and type of disease. The researchers found that the thinning effect was greater the longer people had the disease and some of the medications used to treat bipolar disorder seemed to protect against thinning. The results of this study could be used to evaluate the effectiveness of treatments and aid in early detection and intervention.
Reference:
Cortical abnormalities in bipolar disorder: an MRI analysis of 6503 individuals from the ENIGMA Bipolar Disorder Working Group. Hibar DP, Westlye LT, Doan NT, et al. Mol Psychiatry. 2017 May 2; doi: 10.1038/mp.2017.73.
Making Data FAIR: Findable, Accessible, Interoperable, and Reusable
Members of the data science community recently published a comment article in Scientific Data describing guiding principles for scientific data management and stewardship. Read the article to learn more about this important effort and how the NIH Big Data to Knowledge (BD2K) initiative is contributing to it.
Sustaining the Big-Data Ecosystem
More efficient models are needed for storing, organizing, and accessing biomedical big-data. Read this Nature Outlook perspective (note that full text may require institutional journal access) by the NIH Associate Director for Data Science, Dr. Philip E. Bourne, and the Directors of two NIH Institutes Drs. Lorsch and Green, to learn how the NIH is approaching this issue.
 Dr. Philip Bourne on Biomedical Big Data
Dr. Philip Bourne on Biomedical Big Data
Click on the image to watch a brief video of NIH Associate Data Director Dr. Philip Bourne explaining NIH efforts to coordinate strategies related to computation and informatics in biomedicine across its 27 institutes and centers.
Learn More about the BD2K Initiative and Meet each of its Centers of Excellence:
"NIH Launches a United Ecosystem for Big Data" from Biomedical Computation Review.
NIH Invests Almost $32 Million to Increase Utility of Biomedical Research Data:
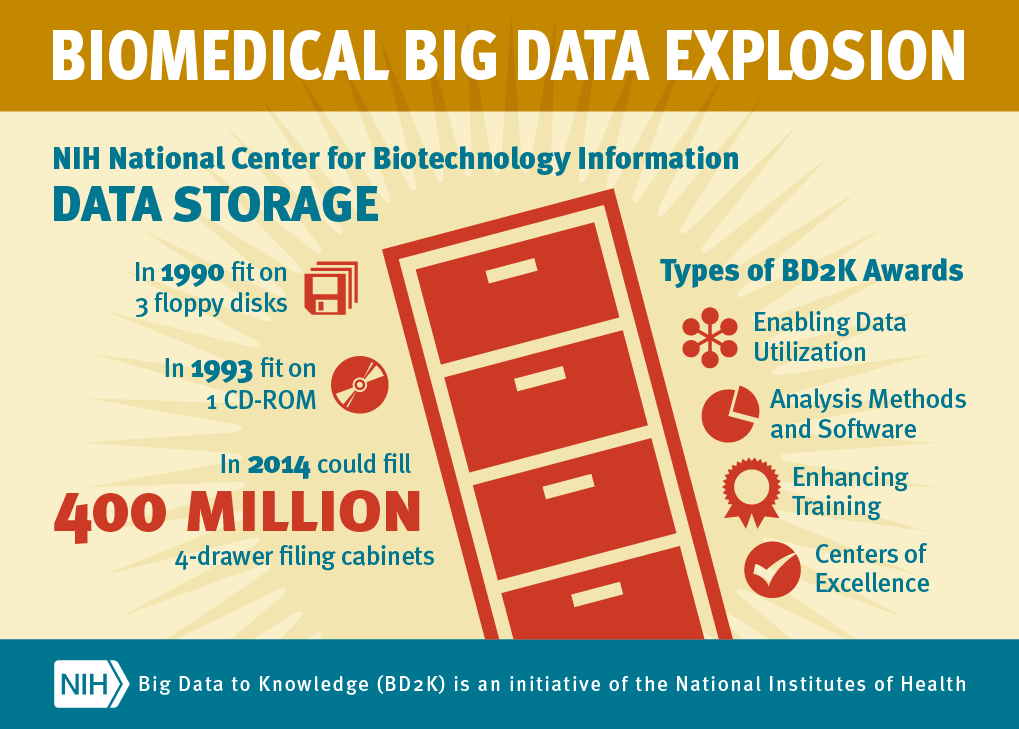 On October 9, 2014, the NIH announced the first grants to be issued through the Big Data to Knowledge initiative. Click on the image at the right to read the Press Release.
On October 9, 2014, the NIH announced the first grants to be issued through the Big Data to Knowledge initiative. Click on the image at the right to read the Press Release.
The NIH Common Fund is contributing support to BD2K efforts involved with:
- Open Educational Resouces for Biomedical Big Data
- Courses for Skills Development in Biomedical Big Data Science
- Mentored Career Development Award in Biomedical Big Data Science for
Clinicians and Doctorally Prepared Students - BD2K-LINCS-Perturbation Data Coordination and Integration Center (DCIC)
- Development of an NIH Data Discovery Index Coordination Consortium
View the Common Fund-supported BD2K research here.
Perspective Paper by BD2K Members Describes the Initiative's Purpose and Goals:
This article by members of the NIH BD2K initiative explains how the NIH is planning to better understand and mine biomedical 'big data' to advance knowledge and promote discovery.



