3D Mapping Strategy Reveals Framework for Gene Expression
Chromatin is a complex of genomic DNA and proteins that make up the chromosomes within the nucleus of a cell. The organization of genomic material into chromatin is presumed to play an important role in regulating expression of genes. However, the precise relationship between spatial genome organization and expression of resident genes in health and disease remains unclear. Toward understanding 3D genome architecture and its relationship to gene regulation, 4D Nucleome researcher Dr. Yijun Ruan, Ph.D., a Jackson Laboratory Professor, Florine Deschenes Roux Chair and Director of Genome Sciences, and his team worked with international collaborators to identify a framework in which genes are organized and transcribed at the chromosomal level.
For these studies, the authors used advanced 3D genome mapping technologies and simulation, as well as super-resolution microscopy. These models revealed higher order chromosome folding and specific chromatin interactions, mediated by the chromatin proteins CTCF and cohesin. These chromatin structures suggest a “barrier” between genes being actively transcribed and those that are not. Importantly, these studies further uncovered potential mechanistic links between genetic mutations associated with specific disease and chromatin topology.
“The significance of this paper lies in our advanced 3D genome mapping strategy,” Dr. Ruan said, “which allowed us to reveal, for the first time, the higher-order and detailed topological structures of the human genome mediated by CTCF and cohesin, and the relation to gene transcription regulation carried out by RNA polymerase II. This publication is also timely adding new excitement to the recently initiated 4D Nucleome program by NIH." For additional information, read The Jackson Laboratory news release.
Reference:
CTCF-Mediated Human 3D Genome Architecture Reveals Chromatin Topology for Transcription. Tang Z, Luo OJ, Li X, Zheng M, Zhu JJ, Szalaj P, Trzaskoma P, Magalska A, Wlodarczyk J, Ruszczycki B, Michalski P, Piecuch E, Wang P, Wang D, Tian SZ, Penrad-Mobayed M, Sachs LM, Ruan X, Wei CL, Liu ET, Wilczynski GM, Plewczynski D, Li G, Ruan Y. Cell. (7) 11611 – 27.
A Novel Approach to Gene Sequencing Reveals Hidden Depths in Microbial Diversity
Advances in DNA sequencing technologies have been a boon for modern human microbiome studies. However, until very recently, these technologies have also had an important limitation. The most common methods have involved the extraction of DNA from these microbiomes and analysis of numerous short stretches of this DNA by sequencing. As the typical microbiome is comprised of thousands of microbial species and millions to trillions of microbial cells, it has been very difficult to re-assemble these short stretches, known as sequence reads, back into the complete genomes of these microbes. Further, with the average bacterial genome about 3,000 base pairs (bp) and the average stretch of DNA sequence read about 100-400 bps, the process of re-assembling millions of these genomes from these short reads has been very difficult.
Reassembling genomes is particularly limiting for performing metagenomics analysis which seeks to uncover the diversity of microbial communities directly from environmental samples, like from the gut tract or skin microbiomes. Finally, although the majority of microbial diversity in microbiomes is found at the subspecies and strain levels, current sequencing technologies have not been able to produce the level of detail needed to get at this level of microbial diversity.
A new study, published December 14, 2015 in Nature Biotechnology, from the laboratory of HMP awardee Dr. Michael Snyder at Stanford University, addresses this important biological problem in the microbiome field with a technical solution. The technique described in Dr. Snyder’s study, used a new sequencing technology, known as TruSeq synthetic long read sequencing technology, to dive deeper into the human gut microbiome. This technology yields 30,000-40,000 bp long reads and allows the investigators to more completely assemble whole microbial genomes from this long read sequence data. More importantly, they were able to consistently recover sufficiently long sequences that allowed them to identify sub-species and strains of bacteria and specific metabolic genes in these strains from these gut microbiome samples and thereby capture the true diversity and metabolic abilities of a microbial community.This now unmasked diversity may lead to new approaches to understanding the specific roles of these microbial strains in human health and disease.
References:
Synthetic long-read sequencing reveals intraspecies diversity in the human microbiome. Kuleshov V, Jiang C, Zhou W, Jahanbani F, Batzoglou S, Snyder M. Nature Biotechnology. 14 December 2015. 10.1038/nbt.316.
Novel Technique Uncovers "Dark" G-Protein Coupled Receptors: Implications for Drug Discovery
 G-protein-coupled receptors (GPCRs) constitute the largest family of proteins encoded by the human genome. With more than 26% of FDA approved drugs acting through them, they are also the most prolific therapeutic targets. However, a large fraction of these receptors are understudied or are considered to be ‘orphan’ receptors and their functions and the chemical signals that control them are not known. Uncovering the biology of these understudied GPCRs could lead to new inroads in therapeutic interventions, which is why the research of Dr. Bryan Roth (University of North Carolina at Chapel Hill) and Dr. Brian Shoichet (University of California, San Francisco) is so exciting.
G-protein-coupled receptors (GPCRs) constitute the largest family of proteins encoded by the human genome. With more than 26% of FDA approved drugs acting through them, they are also the most prolific therapeutic targets. However, a large fraction of these receptors are understudied or are considered to be ‘orphan’ receptors and their functions and the chemical signals that control them are not known. Uncovering the biology of these understudied GPCRs could lead to new inroads in therapeutic interventions, which is why the research of Dr. Bryan Roth (University of North Carolina at Chapel Hill) and Dr. Brian Shoichet (University of California, San Francisco) is so exciting.
These Investigators from the Common Fund’s Illuminating the Druggable Genome program, used an innovative approach to uncover the function of orphan GPCRs. They combined physical and structure-based screens to discover ligands and optimize probes for two under characterized GPCRs –GPR68 and GPR65. GPR68, also known as OGR1, is an orphan receptor that is highly expressed in the brain. The researchers used a newly discovered molecule “ogerin”, which turns on GPR68, to understand the role of this receptor in the brain by monitoring several behaviors in mice. The investigators found that mice that were given ogerin had a dampened response to contextual-based fear memory. The hippocampus is central to this behavior and GPR68 was found to be highly abundant in this brain region. Mice that lacked the GPR68 receptor were not affected by ogerin.
The researchers used the same technique to discover molecules that interacted with another orphan receptor, GPR65. These results not only demonstrate that this approach is applicable to the majority of understudied GPCRs, but it has the potential to be expanded to other classes of proteins as well.
References:
Allosteric ligands for the pharmacologically dark receptors GPR68 and GPR65. Huang XP, Karpiak J, Kroeze WK, Zhu H, Chen X, Moy SS, Saddoris KA, Nikolova VD, Farrell MS, Wang S, Mangano TJ, Deshpande DA, Jiang A, Penn RB, Jin J, Koller BH, Shoichet BK, Roth BL. Nature. 26 November 2015. 527(7579):477-83.
Different Treatments for Crohn’s Disease in Pediatric Patients Have Distinct Effects on the Gut Microbiome
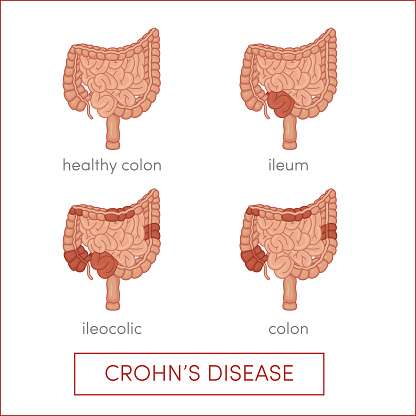
Changes in diet and application of antibiotics and/or anti-inflammatories are the typical interventions used as the standards of care for the treatment of Crohn’s disease (CD), a subtype of inflammatory bowel disease. One major characteristic of CD is an imbalance in the normal composition of the microbiota in comparison to healthy controls. A recent study from Human Microbiome Project awardee Dr. Frederic Bushman and colleagues at the University of Pennsylvania sought to systematically separate the effects of these interventions on the gut microbiomes of a cohort of pediatric CD patients.
Each intervention independently affected the microbiome in CD patients. In particular, antibiotic use seemed to worsen dysbiosis by reducing the abundances of some microbes, increasing the abundances of fungi or both, thus aggravating the condition. Anti-inflammatories, on the other hand, reduced gut microbiota dysbiosis, thereby potentially supporting recovery from CD. Certain defined diets resulted in rapid changes in the gut microbiome suggesting diet may also be an effective treatment for CD.
Since CD patients often have higher rates of gut epithelial cell shedding and/or blood in their stool, stool samples can be sequenced to use as an early indicator of this disease, even before occult blood can be detected. This study suggests that analysis of the microbiome may lead to useful biomarkers for determining the efficacy of standard treatment for CD and for providing additional tests for early detection of CD.
References:
Inflammation, Antibiotics, and Diet as Environmental Stressors of the Gut Microbiome in Pediatric Crohn's Disease. Lewis JD, Chen EZ, Baldassano RN, Otley AR, Griffiths AM, Lee D, Bittinger K, Bailey A, Friedman ES, Hoffmann C, Albenberg L, Sinha R, Compher C, Gilroy E, Nessel L, Grant A, Chehoud C, Li H, Wu GD, Bushman FD. Nature. 14 October 2015. 18(4): 489-500.
The Molecular Libraries and Imaging Program Continues to Pay Dividends After Leaving the Common Fund

The Common Fund’s Molecular Libraries and Imaging (MLI) Program has made promising inroads in the development of new therapeutics for human disease. A compound initially discovered by the NIH Molecular Libraries Probe Production Center at The Scripps Research Institute (TSRI), which was a part of the MLI program, was a precursor to the drug candidate ozanimod; which is currently in two Phase III clinical trials –one for patients with relapsing multiple sclerosis and the other in patients with ulcerative colitis. Dr. Hugh Rosen, a professor at TSRI was a Center Principal Investigator in the MLI program and one of the discoverers of ozanimod. Researchers screened the NIH MLI collection to find allosteric modulators of the sphingosine 1 phosphate receptor 1 (S1P1). Compounds identified in the screen were then optimized for safety and effectiveness in man, resulting in the clinical candidate ozanimod (formerly RPC1063).
Launched in 2004 as one of the original NIH Roadmap programs, the primary intent of MLI was to empower the research community to use small molecule compounds (probes) to study the functions of genes and pathways. The most surprising and encouraging findings, however, have been the great strides MLI is making in drug discovery. The topic of the “Valley of Death” –getting promising scientific breakthroughs made at the bench to the patients, continues to be a high priority area in the biomedical community. The academic approach employed by MLI, in conjunction with the Common Fund’s Structural Biology program, allowed for the exploration of basic biological mechanisms that led to the discovery of a lead compound, the crystal structure of S1P1, and the discovery of the binding site for the compound.
Dr. Rosen is now on the hunt for another drug candidate that has its roots in the MLI program. He will co-direct, as part of the NIH’s Blueprint Neurotherapeutics Network, a project that focuses on a candidate migraine treatment to be tested in preclinical studies. The grant for this project is funded through the National Institute of Neurological Disorders and Stroke.
References:
Celgene bets big on Scripps-originated autoimmunity candidate. Cully M. Nature Reviews Drug Discovery. 21 August 2015. 14(9): 595.
EurekAlert! Press Release from Scripps Research Institute
Common Fund-Supported Researchers Develop Powerful Tools to Edit Genomes

Epigenetic marks are chemical modifications to DNA or DNA-associated proteins that regulate gene expression without changing the DNA sequence. These marks act in many important processes including development, aging, health, and disease; because of this they are targets for therapeutic intervention and intense research activity. Exciting new technologies, such as the CRISPR/Cas9 system used for gene editing (adding, deleting or changing the sequence of targeted genes), have opened new possibilities and sparked a transformation in genetic and epigenomic research (the study of epigenetics across the full genome). Researchers that are funded through the Common Fund’s Epigenomics Program and New Innovator Award at Dr. Gersbach’s lab at Duke University have recently exploited CRISPR/Cas9 technologies to develop state-of-the-art tools that are shaping new discoveries.
Using the powerful CRISPR/Cas9 system, Dr. Gersbach and colleagues made a fusion protein that activates particular genes by directly targeting specific histones for chemical changes, such as acetylation-one type of epigenetic mark. Currently, studying the function of particular epigenetic marks has largely been limited to statistical associations with gene expression patterns, and not to direct functional studies. This unique molecular tool allows researchers to surmount this challenge. With this system, the group showed this directed acetylation to promoter and enhancer regions is sufficient to turn on gene expression. This novel system provides a powerful tool for researchers to directly turn on or off targeted genes of interest thereby allowing direct functional analysis of site-specific epigenetic modifications.
In another innovative application of CRISPR-Cas9 technology, Dr. Gersbach and colleagues were able to control gene expression by simply turning light on or off. They developed a system where they made two fusion proteins, fusion proteins are made by the joining of two or more genes that originally coded for separate proteins. One protein consisted of an inactivated Cas9 protein fused with a plant protein called CIB1 and the other protein fused a transcriptional activation domain to cryptochrome 2 (CRY2). When both proteins and a guide RNA are present in cells and illuminated with blue light, the two fusion proteins pair up, bind to their DNA, and turn on gene expression. Future application of this innovative technique could allow researchers to target any gene in an organism and turn it on or off with the switch of a light, which has amazing potential to transform genetic engineering.
References:
Epigenome Editing by a CRISPR-Cas9-Based Acetyltransferase Activates Genes from Promoters and Enhancers. Hilton, I. B., A. M. D'Ippolito, C. M. Vockley, P. I. Thakore, G. E. Crawford, T. E. Reddy, and C. A. Gersbach. Nature Biotechnology. 33(5): 510-517.
A Light-Inducible CRISPR-Cas9 System for Control Of Endogenous Gene Activation. Polstein, L. R., and C. A. Gersbach. Nature Chemical Biology. 11(3): 198-200.
NIH-Funded KOMP2 Program Leads the Way in Efforts to Ensure Reproducibility and Enhance Transparency in Research
Researchers funded by the National Institutes of Health’s Common Fund Knockout Mouse Phenotyping Program (KOMP2) are making efforts to improve methods of data collection and reporting in animal research, with the goals of making these processes more accessible and research findings more reproducible. The KOMP2 program, a part of the International Mouse Phenotyping Consortium (IMPC), aims to extensively test and generate data about mice with disrupted, or “knocked out,” genes to help better understand human biology and disease. KOMP2 researchers are examining the clinical and physical characteristics – phenotypes – of mice carrying mutations in approximately 20,000 specific genes across the genome. They hope to systematically describe each gene and its biological function.
In 2010, the United Kingdom National Centre for the Replacement, Refinement, and Reduction of Animals in Research (3R) introduced the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines . These guidelines were designed to address an emerging problem in biomedical animal studies -- a lack of reproducibility of research results -- and improve the overall communication of research findings. The ARRIVE guidelines consist of a checklist to be used when submitting manuscripts that include animal research. They lay out reporting requirements to ensure all the information is available to allow fully reproducible research.
To enhance and embrace reproducibility in the IMPC large-scale research data collection and analyses efforts, KOMP2-funded investigators modified the ARRIVE guidelines to apply them to the large international database housing all the program data . The investigators described the process and challenges of applying the guidelines to the IMPC database May 20, 2015, in PLoS Biology. Their efforts assessed and documented how each of the IMPC centers carried out experimental procedures; details on how the data were obtained are available for download from the IMPC web portal. Investigators also developed a new and standardized language for the IMPC consortium, allowing all phenotypic data to be described in the same way and compared across research sites.
Reference:
Applying the ARRIVE Guidelines to an In Vivo Database. Karp N, Meehan T, Morgan H, Mason J, Blake A, Kurbatova N, Smedley D, Jacobsen J, Mott R, Iyer V, Matthews P, Melvin D, Wells S, Flenniken A, Masuya H, Wakana S, White J, Lloyd KC, Reynolds C, Paylor R, West D, Svenson K, Chesler E, de Angelis M, Tocchini-Valentini G, Sorg T, Herault Y, Parkinson H, Mallon A, Brown S. PLOS Biology. 20 May 2015.
Financial Incentives can Increase the Effectiveness of Smoking Cessation Programs

Smoking remains the leading cause of preventable illness and death despite the implementation of national policies, behavioral programs, and pharmacological treatments that have helped decrease smoking rates in the US. One promising line of research in smoking cessation relies on findings from the field of behavioral economics, which applies what we know about how people make health-related decisions to the design of health-behavior change interventions. Often, interventions informed by behavioral economics make use of financial incentives to encourage healthy behaviors and promote behavior change. Science of Behavior Change awardee Dr. Scott Halpern and colleagues at the University of Pennsylvania report their findings that financial incentive programs can lead to sustained cessation of smoking, and that the structures of incentive programs are differentially efficacious. Moreover, these incentive programs were more effective than the usual method of care, which included informational resources and free smoking cessation aids.
Study participants included CVS/Caremark employees or their relatives from across the US. After enrolling through a web-based research portal, participants were randomized into one of four incentive programs: individual reward, individual deposit, collaborative reward and collaborative deposit. The reward conditions differed by whether monetary rewards were paid based on individual or group success in smoking cessation, and by the initial, personal or collaborative monetary investment required of participants. The deposit conditions required individuals or groups to deposit some of their own money into an account that was enhanced by matched funds; they would get their own money back plus the additional funds if they successfully quit smoking, but lose their deposit and the additional funds if they were unsuccessful. The reward conditions did not require an initial investment of personal or collaborative funds, and so did not involve risk of loss of any participant’s own funds. Participants could opt out of assignment to any of the conditions, in which case they were re-assigned to another condition in the study. The logic behind the deposit conditions is that behavioral economics has shown that people are very motivated to avoid losses of their own funds; much more motivated than they are to gain rewards of the same or even larger size.
Researchers found that all four incentive programs were effective in promoting sustained smoking abstinence. Perhaps not surprisingly, fewer participants accepted conditions that required them to deposit their own money as part of their reward. Those who did accept the deposit condition, however, were substantially more successful in quitting smoking than those who accepted conditions that did not require an initial deposit. Interestingly, although the collaborative reward conditions have sometimes been found to be more efficacious in spurring behavior change in the past, this was not the case in this study.
Over all, this study underlined the fact that incentives-based programs can be very effective in helping people quit smoking, but the level of effectiveness depends on both the acceptability of intervention and structure of the incentive intervention. The deposit-based incentive intervention was far more efficacious for those who accepted it, but far less palatable to participants who were offered it. Future work in the area might target ways in which the acceptability of the more efficacious interventions could be increased, as well as increasing the efficaciousness of the interventions people are more likely to accept.
References:
A randomized trial of four financial incentive programs for smoking cessation. Halpern SD, French B, Small DS, Saulsgiver KA, Harhay MO, Audrain-McGovern J, Loewenstein G, Brennan TA, Asch DA, Volpp KG. N Engl J Med. 2015
Read the Press Release from Perelman School of Medicine at the University of Pennsylvania.
New Resource Shows How Differences in DNA Affect Gene Activity and Disease Susceptibility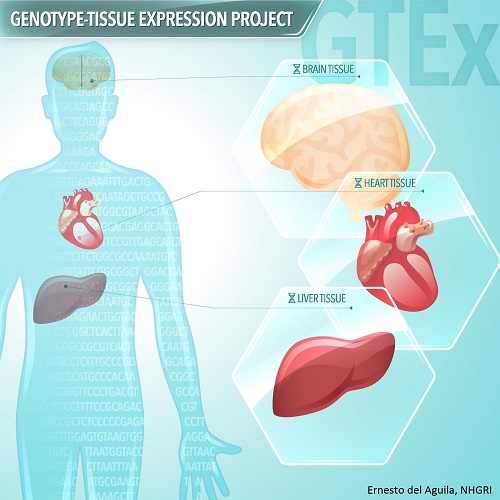
We have known for many years that differences in the DNA that codes for our genes affect everything from our eye color to our susceptibility for certain diseases. Now we are finding that differences in genes are only part of the story. Differences in DNA that control when and how much genes are turned on and off can have a profound impact on health and disease. Researchers funded by the Genotype-Tissue Expression (GTEx) program have created a new data resource to help find out how differences in an individual’s genetic make-up can affect gene activity and contribute to disease.
Scientists can use the new resource to examine genes and gene regulation in many different types of human tissues at the same time. Investigators are collecting more than 30 tissue types from autopsy or organ donations and tissue transplant programs, and analyzing both DNA and RNA from samples. The project will eventually include tissue samples from about 900 deceased donors.
The resource is already beginning to bear fruit. One study looked at mutations called protein-truncating variants which shorten the protein-coding sequence of genes. Rare protein-truncating variants can lead to diseases like Duchenne muscular dystrophy. Most protein-truncating variants are harmless, and researchers found that each person’s genome carries about 100 of them. Another study looked at gene activity in a variety of tissues from multiple donors. Researchers found slightly fewer than 2,000 genes that vary with age, including genes related to Alzheimer’s disease. They also found more than 750 genes with differences in activity between men and women, with most in breast tissue.
References:
The Genotype-Tissue Expression (GTEx) Pilot Analysis: Mutitissue Gene Regulation in Humans. The GTEx Consortium. Science, May 2015, Vol. 348 no. 6235 pp. 648-660. Read the article abstract.
Read the NIH Press Release.
Read more about the Genotype-Tissue Expression (GTEx) program here.
A list of companion papers to the GTEx Pilot Analysis is available via the GTEx Portal website.
New method using llamas is set to transform biomedical research
Llamas have become the surprising focal point of a new technology that is aimed at making “nanobodies” more accessible to the research community. Nanobodies, considered to be the “little cousins” of antibodies, are therapeutic proteins derived from their larger antibody cousins. Antibodies are proteins produced by the immune system and used to identify and neutralize foreign elements in the body. They are extremely valuable research tools that are used routinely in basic research, diagnostic testing and therapeutics. Nanobodies can perform the same functions as antibodies; however, their small size and extreme stability make them attractive replacements. Although the appeal of using nanobodies in research has been clear for some time, the difficulties of producing them in sufficient quantities have limited their usefulness.
A discovery in llamas has changed the trajectory of nanobody production. Although not the original developers of nanobodies, a research team headed by Michael Rout, an investigator funded by the National Center for Dynamic Interactome Research http://www.ncdir.org/, within the NIH Common Fund’s National Technology Centers for Networks and Pathways (TCNPs) http://commonfund.nih.gov/bbpn/overview#TCNPs initiative has developed a novel approach to increasing nanobody production. Dr. Rout and his colleagues have designed a faster more direct technique to generate nanobodies, making them more accessible to the scientific community. They chose the llama instead of more common laboratory animals as a model for this method because llamas were found to make a unique subset of antibodies that can easily be turned into nanobodies. Faster production of these highly versatile nanobodies has the potential to transform research and catalyze scientific advances in a timeframe of weeks to months, instead of years.
Reference:
A robust pipeline for rapid production of versatile nanobody repertoires. Fridy PC, Li Y, Keegan S, Thompson MK, Nudelman I, Scheid JF, Oeffinger M, Nussenzweig MC, Fenyö D, Chait BT, Rout MP. Nat Methods. 2014 Dec;11(12):1253-60.
Press Release: New technique efficiently turns antibodies into highly tuned ‘nanobodies’
Common Fund-supported researchers map epigenome of more than 100 tissue and cell types
More than 440 researchers in 32 labs around the world participating in the Common Fund’s Roadmap Epigenomics program have generated and analyzed 111 reference human epigenomes. This is the largest collection to date of reference human epigenomes from a broad range of representative primary cells and tissues. A genome is defined as the DNA sequences present in a cell, and an epigenome refers to the chemical modifications and non-sequence changes to DNA and DNA-associated proteins. Currently, researchers are beginning to understand the many consequences of these chemical modifications, and early insights from this new analysis reveal that different epigenomic states are associated with differences in age, sex, and tissue type. The many companion papers describing additional analyses demonstrate the broad applicability of the resource. These studies examine the biological importance of epigenetic changes in the context of stem cell differentiation, obesity, Alzheimer’s disease, cancer, and cardiac disease. This remarkable accomplishment will usher in new scientific understanding of the complexity of the human genome at a deeper level and spur new research advances in human health and disease.
Reference:
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature, Feb. 18, 2015; 518; 317-330.
Read the NIH press release here.
Read more about the Epigenomics program here.
See video on the Epigenome by Nature here.
Study Finds Potential New Drug Target for Lung Cancer
 Scientists have long known that the metabolism of tumor cells differs from normal, healthy cells. However, it has been challenging to study tumor metabolism in living tumor cells from a large number of cancer patients. Metabolomics, a technique that allows scientists to analyze the molecules involved in metabolism, may change that. Researchers at the Common Fund-supported Resource Center for Stable Isotope-Resolved Metabolomics (RC-SIRM) looked at how the molecule glucose is broken down as part of the metabolism of patients with non-small-cell lung cancer. Shortly before surgery for tumor removal, the researchers injected patients with a form of glucose that is non-radioactively labeled. Using metabolomic techniques, researchers can follow how the labeled glucose is broken down by both the tumor cells and healthy cells from the same patient. As the glucose is broken down into component molecules, the label will remain with one or more of the components. These labeled components are a clue to which metabolic processes are active in the tumor.
Scientists have long known that the metabolism of tumor cells differs from normal, healthy cells. However, it has been challenging to study tumor metabolism in living tumor cells from a large number of cancer patients. Metabolomics, a technique that allows scientists to analyze the molecules involved in metabolism, may change that. Researchers at the Common Fund-supported Resource Center for Stable Isotope-Resolved Metabolomics (RC-SIRM) looked at how the molecule glucose is broken down as part of the metabolism of patients with non-small-cell lung cancer. Shortly before surgery for tumor removal, the researchers injected patients with a form of glucose that is non-radioactively labeled. Using metabolomic techniques, researchers can follow how the labeled glucose is broken down by both the tumor cells and healthy cells from the same patient. As the glucose is broken down into component molecules, the label will remain with one or more of the components. These labeled components are a clue to which metabolic processes are active in the tumor.
The RC-SIRM team found that tumor cells broke down the glucose using two processes, glycolysis and the tricarboxylic acid cycle (TCA cycle). They also found that an enzyme involved in the TCA cycle, called pyruvate carboxylase, was produced in greater quantities in tumors compared to healthy tissue. Reducing the level of this enzyme in lab-grown human lung cancer cells reduced cell growth and interfered with the ability of these cells to form tumors in mice. This study by the RC-SIRM team demonstrates the exciting potential for metabolomic approaches to reveal new pathways and players in cancer metabolism that may become novel targets for cancer therapy and diagnostics.
RC-SIRM at the University of Kentucky is a Regional Metabolomics Resource Core funded through the Common Fund Metabolomics program. Learn more about the Metabolomics program.
Read the press release and watch the video from the University of Kentucky on this discovery.
The original research article was published in the Journal of Clinical Investigation. Read the article abstract.
Protein Tangles Spell Trouble for Cancer Cells
New research has demonstrated that a destructive biological process involved in Alzheimer’s disease may  hold promise as a novel treatment strategy for cancer. Anomalies in protein folding can generate harmful protein deposits within cells, called amyloids. These amyloid deposits underlie several neurodegenerative disorders in humans, particularly Alzheimer's disease. While amyloids are widely recognized as killers of neurons, a new study from New Innovator Awardee Dr. Chengkai Dai at The Jackson Laboratory reveals that cancer cells are also vulnerable to these toxic proteins. Cancer cells manage to contain deleterious amyloids by hijacking a cellular stress response that has evolved to counter protein misfolding, thereby sustaining the cancer cells’ vicious survival and growth. But importantly, this study also points to a possible means of countering this effect. Surprisingly, the authors’ findings reveal that cancer cells, unlike normal cells, constantly suffer heightened intrinsic stress. As a result, cancer cells are therefore particularly susceptible to extrinsic insults, including the activity of therapeutic agents that either perturb protein folding or subdue the defensive stress response. These agents efficiently induce the generation of amyloids within cancer cells and thereby exert potent anti-cancer effects. Thus, this study highlights a potential therapeutic strategy that harnesses the culprit behind neurodegenerative diseases to combat malignancy.
hold promise as a novel treatment strategy for cancer. Anomalies in protein folding can generate harmful protein deposits within cells, called amyloids. These amyloid deposits underlie several neurodegenerative disorders in humans, particularly Alzheimer's disease. While amyloids are widely recognized as killers of neurons, a new study from New Innovator Awardee Dr. Chengkai Dai at The Jackson Laboratory reveals that cancer cells are also vulnerable to these toxic proteins. Cancer cells manage to contain deleterious amyloids by hijacking a cellular stress response that has evolved to counter protein misfolding, thereby sustaining the cancer cells’ vicious survival and growth. But importantly, this study also points to a possible means of countering this effect. Surprisingly, the authors’ findings reveal that cancer cells, unlike normal cells, constantly suffer heightened intrinsic stress. As a result, cancer cells are therefore particularly susceptible to extrinsic insults, including the activity of therapeutic agents that either perturb protein folding or subdue the defensive stress response. These agents efficiently induce the generation of amyloids within cancer cells and thereby exert potent anti-cancer effects. Thus, this study highlights a potential therapeutic strategy that harnesses the culprit behind neurodegenerative diseases to combat malignancy.
Reference:
Tang et al. MEK guards proteome stability and inhibits tumor-suppressive amyloidogenesis via HSF-1. Cell, 160, Feb 2015.
New Device Holds Promise for Hard-to-Reach Tumors
Pioneer awardee Dr. Joseph DeSimone has developed a device that uses electrical currents to drive  chemotherapy drugs directly into tumors, potentially revolutionizing how doctors treat some of the most challenging types of cancer. Several types of cancer, notably pancreatic cancer, can be difficult to treat with surgery because the tumor can become enmeshed with healthy tissue and blood vessels. Intravenous chemotherapy treatment may also be ineffective, because these tumors often don’t have a good enough blood supply to deliver effective concentrations of the drug. To overcome these obstacles, Dr. DeSimone and colleagues at the University of North Carolina Chapel Hill, Duke University, and North Carolina State University designed a device that can be implanted directly on the tumor or applied to the skin to precisely deliver therapeutic doses chemotherapy drugs into the tumor, with minimal amounts of the drug spreading to other places in the body. In animal models of pancreatic and breast cancer, treatment with the device slowed tumor growth or even reduced tumor size. Treatment with the device was even more effective when combined with standard intravenous chemotherapy and radiation. These results suggest that the new approach pioneered by Dr. DeSimone may one day be added to the arsenal of weapons doctors use in the fight against cancer.
chemotherapy drugs directly into tumors, potentially revolutionizing how doctors treat some of the most challenging types of cancer. Several types of cancer, notably pancreatic cancer, can be difficult to treat with surgery because the tumor can become enmeshed with healthy tissue and blood vessels. Intravenous chemotherapy treatment may also be ineffective, because these tumors often don’t have a good enough blood supply to deliver effective concentrations of the drug. To overcome these obstacles, Dr. DeSimone and colleagues at the University of North Carolina Chapel Hill, Duke University, and North Carolina State University designed a device that can be implanted directly on the tumor or applied to the skin to precisely deliver therapeutic doses chemotherapy drugs into the tumor, with minimal amounts of the drug spreading to other places in the body. In animal models of pancreatic and breast cancer, treatment with the device slowed tumor growth or even reduced tumor size. Treatment with the device was even more effective when combined with standard intravenous chemotherapy and radiation. These results suggest that the new approach pioneered by Dr. DeSimone may one day be added to the arsenal of weapons doctors use in the fight against cancer.
Read the press release from UNC Chapel Hill
Read the NBC News story "Device Reaches Where Cancer Drugs Cannot"
Reference:
Byrne et al. Local ionophoretic administration of cytotoxic therapies to solid tumors. Science Translational Medicine, Feb 2015, 7(273).


