Science Highlights Archives
Better Biomarkers for Muscular Dystrophy
 New and non-invasive methods to identify muscular dystrophy (MD) are greatly needed to replace painful and highly invasive muscle biopsy techniques. Currently, clinicians and scientists typically isolate an important biological molecule called RNA from muscle cells removed during a biopsy to look for RNA that has not been properly processed through a process called splicing. The mis-spliced RNA can serve as a diagnostic biomarker for MD. However, RNA doesn't have to be obtained only from cells taken from muscle biopsy. In some cases, RNA found outside of cells called extracellular RNA (exRNA), can be isolated from bodily fluids like urine and serve as biomarkers for some diseases. Researchers from the NIH Common Fund’s ExRNA Communication program examined if exRNA in urine could be used to detect the mis-spliced RNA from muscular dystrophy patients.
New and non-invasive methods to identify muscular dystrophy (MD) are greatly needed to replace painful and highly invasive muscle biopsy techniques. Currently, clinicians and scientists typically isolate an important biological molecule called RNA from muscle cells removed during a biopsy to look for RNA that has not been properly processed through a process called splicing. The mis-spliced RNA can serve as a diagnostic biomarker for MD. However, RNA doesn't have to be obtained only from cells taken from muscle biopsy. In some cases, RNA found outside of cells called extracellular RNA (exRNA), can be isolated from bodily fluids like urine and serve as biomarkers for some diseases. Researchers from the NIH Common Fund’s ExRNA Communication program examined if exRNA in urine could be used to detect the mis-spliced RNA from muscular dystrophy patients.
In this study, researchers isolated exRNA from the urine of patients with a certain type of Muscular Dystrophy (myotonic dystrophy type I (DM1)) and from healthy volunteers. They found several different RNA molecules that were not spliced properly in DM1 patients. When 10 of the mis-spliced RNA molecules were all present in one patient, the researchers could predict with 100% accuracy that the people had the disease. The presence of these 10 mis-spliced RNA molecules is the composite biomarker signature the researchers were hoping to identify. While more studies are needed, this approach is a promising alternative to invasive muscle biopsies. The exRNA approach also showed promise in detecting differences in RNA splicing in patients with DM1 who were being treated with drugs designed to target the RNA splicing process. This may also represent a painless and low-cost way to track disease activity or response to drug treatment.
Reference:
Analysis of extracellular mRNA in human urine reveals splice variant biomarkers of muscular dystrophies. Antoury L, Hu N, Balaj L, Das S, Georghiou S, Darras B, Clark T, Breakefield XO, Wheeler TM. Nature Communications. 2018 Sep 25;9(1):3906.
In the News: Extracellular RNA in urine may provide useful biomarkers for muscular dystrophy. Mass General News Release. September 25, 2018.
New RNA Nanoparticle Helps Deliver RNA to Fight Cancer
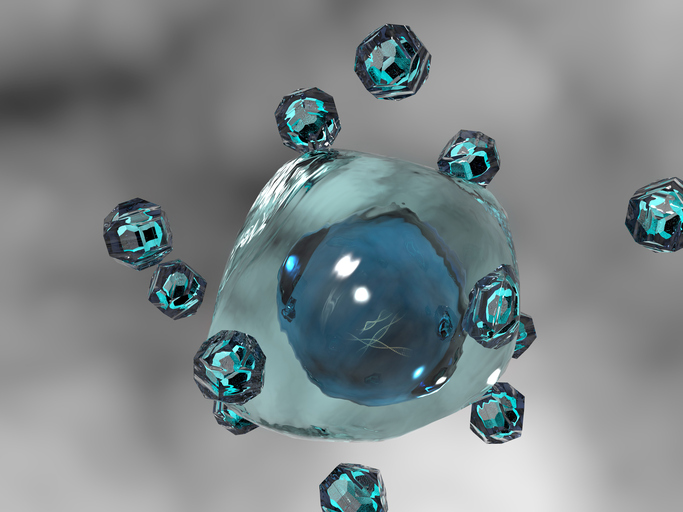 Attaching RNA nanoparticles to containers that shuttle back forth between cells may help deliver therapeutics to destroy cancer cells. RNAs are key molecules in many cellular processes, ranging from gene regulation to protein synthesis to signaling between cells. Recent research has also used RNA as structural decoys to look like other normal parts of cells, such as binding motifs or gene regulators. These “RNA nanoparticles” are part of promising efforts to fight cancer development and progression. However, several major hurdles remain before this becomes a clinical reality; one challenge is targeting the right cells and another is escaping natural defense mechanisms. The following study from the Extracellular RNA Communication Program, brings us one step closer to overcoming these challenges and unlocking the transformative potential of this new area of research for human health, disease diagnosis, and treatment.
Attaching RNA nanoparticles to containers that shuttle back forth between cells may help deliver therapeutics to destroy cancer cells. RNAs are key molecules in many cellular processes, ranging from gene regulation to protein synthesis to signaling between cells. Recent research has also used RNA as structural decoys to look like other normal parts of cells, such as binding motifs or gene regulators. These “RNA nanoparticles” are part of promising efforts to fight cancer development and progression. However, several major hurdles remain before this becomes a clinical reality; one challenge is targeting the right cells and another is escaping natural defense mechanisms. The following study from the Extracellular RNA Communication Program, brings us one step closer to overcoming these challenges and unlocking the transformative potential of this new area of research for human health, disease diagnosis, and treatment.
Using new nanotechnologies, researchers decorated the outside of extracellular vesicles (EVs) – containers that shuttle back and forth between cells – with RNA nanoparticles made to look like a normal antibody. This provided an effective signal to find and bind certain cancer cells. The EVs were engineered to contain another RNA that was able to get inside the cancer cells, bind to specific DNA targets, and stop cells from dividing and making more cancer cells. Most importantly, this delivery system was able to ensure that the RNA avoided “endosome traps,” – part of natural defense mechanisms that can destroy the RNA before it has a chance to act. While this study used cells grown in the lab and animal models, it offers a promising roadmap for future clinical studies.
Reference:
Nanoparticle orientation to control RNA loading and ligand display on extracellular vesicles for cancer regression. Binzel, P. F., et al. Nature Nanotechnology. 2017 Dec 11. doi: 10.1038/s41565-017-0012-z.
New Study Offers Promise for the Safety of RNA Therapeutics
 Researchers supported by the Common Fund’s Extracellular RNA Communication (ERC) program at The Ohio State University are studying the safety of delivering RNA in extracellular vesicles (EVs) as a possible disease treatment. RNA is a biological molecule that makes protein. Proteins perform a variety of essential functions, from providing structure to our bodies to protecting us from bacteria. RNA is found primarily inside cells and only recently has been found outside of cells and considered as a potential treatment option. For example, some recent evidence suggested that RNAs can be developed as therapeutics for cancer or multiple sclerosis. The natural ability of EVs to transfer biologic materials like RNA throughout the human body is an exciting system to harness. Current efforts, including those of the ERC program, are exploring whether EVs containing RNA could be effective therapeutics or if EVs can be vehicles for drug delivery to treat diseases like cancers or brain disorders. Scientists are also working to understand the potential harmful side effects or unintended immune complications of EVs and their RNA cargo. While proving a treatment has a desired effect is critical in early preclinical studies, it can be even more critical to know if a potential treatment could have toxic side effects or may elicit a dangerous immune response.
Researchers supported by the Common Fund’s Extracellular RNA Communication (ERC) program at The Ohio State University are studying the safety of delivering RNA in extracellular vesicles (EVs) as a possible disease treatment. RNA is a biological molecule that makes protein. Proteins perform a variety of essential functions, from providing structure to our bodies to protecting us from bacteria. RNA is found primarily inside cells and only recently has been found outside of cells and considered as a potential treatment option. For example, some recent evidence suggested that RNAs can be developed as therapeutics for cancer or multiple sclerosis. The natural ability of EVs to transfer biologic materials like RNA throughout the human body is an exciting system to harness. Current efforts, including those of the ERC program, are exploring whether EVs containing RNA could be effective therapeutics or if EVs can be vehicles for drug delivery to treat diseases like cancers or brain disorders. Scientists are also working to understand the potential harmful side effects or unintended immune complications of EVs and their RNA cargo. While proving a treatment has a desired effect is critical in early preclinical studies, it can be even more critical to know if a potential treatment could have toxic side effects or may elicit a dangerous immune response.
In this study, researchers form the ERC program used normal cells, which naturally produce EVs, as well as cells engineered to make EVs carrying a specific type of RNA inside them. They treated mice with EVs collected from both cell types for a three-week period and carefully monitored if normal EVs or engineered EVs caused any toxic or immune responses. They used multiple monitoring techniques, including visual examination of mice, measuring blood chemistry profiles, dissecting organs, and measuring immune markers. Overall, these assays they found no potential harm from the EVs of normal and engineered cells, though small changes in immune responses may require further study. Next steps will determine if higher doses of EVs or a longer dosing period would be as safe, if EVs from different cell types act differently, and if EVs from human cells are immunogenic or toxic in other animal models. This study shows early promising signs for the safe use of EVs containing RNAs, and presents a standard framework for comprehensive study of immune effects and toxicity for future studies.
Reference:
Comprehensive toxicity and immunogenicity studies reveal minimal effects in mice following sustained dosing of extracellular vesicles derived from HEK293T cells. X. Zhu, M. Badawi, S. Pomeroy, D.S. Sutaria, Z. Xie, A. Baek, J. Jiang, O. A. Elgamal, X. Mo, K. La Perle, J. Chalmers, T. D. Schmittgen, and M.A. Phelps. Journal of Extracellular Vesicles. Vol. 6, Iss. 1,2017.
Combining Anti-RNA Treatments With Chemotherapy May Offer Benefits for Lung Cancer Patients
 Chemotherapy is a standard treatment for many cancers. However, resistance to chemotherapy can develop over time, which causes cancer cells to lose the ability to respond effectively to treatment. It is unknown how this resistance develops. Researchers from the Common Fund’s Extracellular RNA Communication program have begun to examine the role extracellular RNAs may play in this resistance by studying a particular RNA called miR-155. MiR-155 is found at increased levels in many cancers. In forms of cancer that are particularly aggressive and resistant to treatment, miR-155 is found at even higher levels. The potential role of miR-155 in cancer and resistance to chemotherapy is not well-understood. Does miR-155 influence resistance to chemotherapy? Can available drugs targeted against miR-155 (“anti-miR-155”) be used to stop or reverse chemotherapy resistance? To address these questions, researchers explored how miR-155 works in lung cancer and evaluated if a treatment to inhibit miR-155 could help stop resistance to chemotherapy.
Chemotherapy is a standard treatment for many cancers. However, resistance to chemotherapy can develop over time, which causes cancer cells to lose the ability to respond effectively to treatment. It is unknown how this resistance develops. Researchers from the Common Fund’s Extracellular RNA Communication program have begun to examine the role extracellular RNAs may play in this resistance by studying a particular RNA called miR-155. MiR-155 is found at increased levels in many cancers. In forms of cancer that are particularly aggressive and resistant to treatment, miR-155 is found at even higher levels. The potential role of miR-155 in cancer and resistance to chemotherapy is not well-understood. Does miR-155 influence resistance to chemotherapy? Can available drugs targeted against miR-155 (“anti-miR-155”) be used to stop or reverse chemotherapy resistance? To address these questions, researchers explored how miR-155 works in lung cancer and evaluated if a treatment to inhibit miR-155 could help stop resistance to chemotherapy.
Interestingly, they showed that increased levels of miR-155 induce resistance to many different types of chemotherapeutic agents. They also demonstrated that miR-155 partners with TP53, a well-known tumor suppressor, to promote resistance. Furthermore, when they added a drug that blocks miR-155, they found resistance lessened. This means that even though resistance developed, it can be reversed and the chemotherapeutic drugs can become effective at killing cancerous cells again. Excitingly, this anti-miR-155 treatment had no toxic side effects in mice, suggesting the treatment has potential to be used in clinical trials in combination with standard chemotherapy regimens.
Reference:
Combining anti-miR-155 with chemotherapy for the treatment of lung cancers. Van Roosbroeck, K., F. Fanini, T. Setoyama, C. Ivan, C. Rodriguez-Aguayo, E. Fuentes-Mattei, L. Xiao, I. Vannini, R. Redis, L. D'Abundo, X. Zhang, M. S. Nicoloso, S. Rossi, V. Gonzalez-Villasana, R. Rupaimoole, M. Ferracin, F. Morabito, A. Neri, P. Ruvolo, V. R. Ruvolo, C. V. Pecot, D. Amadori, L. Aruzzo, S. Calin, X. Wang, M. J. You, A. Ferrajoli, R. Z. Orlowski, W. Plunkett, T. Lichtenberg, R. V. Davuluri, I. Berindan-Neagoe, M. Negrini, Wistuba, II, K. Hagop, A. K. Sood, G. Lopez-Berestein, M. J. Keating, M. Fabbri and G. A. Calin. Clinical Cancer Research.2017 Jun 1; 23(11): 2891–2904.
For more information see the exRNA Consortium blog!
Circular RNAs May Offer Early Signs of Colorectal Cancer
 Researchers supported by the Common Fund’s Extracellular RNA Communication (ERC) program have found that circular RNAs (circRNA) may one day serve as promising biomarkers for colorectal cancer. Currently, not much is known about circular RNAs, which are RNAs that don’t code for a gene and are found in a circular conformation, rather than the more well-known linear conformation. Until very recently, circular RNAs were often thought to be mistakes created by normal biological processes and with no known function. Furthermore, what these circRNAs are doing when released outside of cells has not been well explored.
Researchers supported by the Common Fund’s Extracellular RNA Communication (ERC) program have found that circular RNAs (circRNA) may one day serve as promising biomarkers for colorectal cancer. Currently, not much is known about circular RNAs, which are RNAs that don’t code for a gene and are found in a circular conformation, rather than the more well-known linear conformation. Until very recently, circular RNAs were often thought to be mistakes created by normal biological processes and with no known function. Furthermore, what these circRNAs are doing when released outside of cells has not been well explored.
The ERC program researchers explored the presence of circRNAs inside cells and the release of circRNAs outside cells in colorectal cancer models, and discovered several surprising findings. First, colorectal cancer cells with mutations in the KRAS oncogene had levels of circRNAs that were much lower compared to normal cells. This was not dependent on changes in the cell’s DNA that codes for the RNA, suggesting that KRAS acts by a global mechanism influencing overall circRNA levels. Because KRAS is an oncogene, its influence on levels of circRNAs suggests that circRNAs may be involved in oncogenesis, or the development of cancer. Second, circRNAs could also be detected in exosomes, which are small vesicles secreted from cells. Furthermore, some selected circRNAs were at higher levels, while others were at lower levels in exosomes released from mutant cell lines. This suggests that researchers may one day be able to test for particular circRNAs in exosomes to diagnose colorectal cancer. However, more research is needed to determine how exactly circRNAs are regulated and get into exosomes, and if they play a role in the development or spread of cancer.
Reference:
Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Dou, Y., Cha, D. J., Franklin, J. L., Higginbotham, J. N., Jeppesen, D. K., Weaver, A. M., Prasad, P., Levy, S., Coffey, R.J., Patton, J.G., and Zhang, B. Scientific Reports. 2016; (6)37982:1-11.
For more information see the exRNA Consortium blog!
Complex and Diverse RNA Found in Human Plasma
 Until recently, most studies of extracellular RNA (exRNA) in human body fluids have investigated samples from only a handful of individuals and focused mainly on a single type of exRNA - microRNA. Because of this, there is uncertainty about the amount and diversity of other types of exRNAs in large and healthy populations. Recently, researchers in the ExRNA Communication Program collaborated to measure exRNAs in the blood plasma of over 2700 participants from the Framingham Heart Study, a well-known large observational cohort study based in Framingham, MA. They characterized almost 500 of the most abundant exRNA species. Surprisingly, many non-human exRNAs were also found in the plasma samples. While this finding is similar to reports from other labs, how these non-human molecules, potentially from food sources, work and get into in the plasma is not currently known.
Until recently, most studies of extracellular RNA (exRNA) in human body fluids have investigated samples from only a handful of individuals and focused mainly on a single type of exRNA - microRNA. Because of this, there is uncertainty about the amount and diversity of other types of exRNAs in large and healthy populations. Recently, researchers in the ExRNA Communication Program collaborated to measure exRNAs in the blood plasma of over 2700 participants from the Framingham Heart Study, a well-known large observational cohort study based in Framingham, MA. They characterized almost 500 of the most abundant exRNA species. Surprisingly, many non-human exRNAs were also found in the plasma samples. While this finding is similar to reports from other labs, how these non-human molecules, potentially from food sources, work and get into in the plasma is not currently known.
This is the first study to report widespread and variable levels of many different types of small RNAs in human plasma. The significance of this newly discovered diversity of exRNAs for biological processes or disease is uncertain. Future work will determine how the presence of these exRNAs in the circulation may influence normal human processes or diseases. Understanding the origin and possible functions of these exRNAs will be a significant and necessary step before exRNAs can be used in the clinic as therapeutic molecules or biomarkers of disease.
Reference:
Diverse human extracellular RNAs are widely detected in human plasma. Freedman JE, Gerstein M, Mick E, Rozowsky J, Levy D, Kitchen R, Das S, Shah R, Danielson K, Beaulieu L, Navarro FC., Nature Communications. 2016 Apr 26;7;11106. doi: 10-1038/ncomms11106.
Uncovering Key Pathways and Players in Extracellular RNA Sorting
 RNA carried in Extracellular Vehicles (EVs) and transported to neighboring or distant cells in the body can influence gene expression and other cellular process. However, not much is known about the mechanisms involved in the underlying processes. For example, how are EVs generated that carry these RNAs, and how are different types of RNAs, like microRNA (miRNA), sorted into different types of EVs? In this work, researchers shed light on some of the basic mechanisms and regulatory signals involved in sorting and transport of extracellular RNAs. Using well-characterized cancer cell lines, the researchers demonstrate that an RNA binding protein called Ago2, a component of the RNA-induced silencing complex (RISC), is responsible for helping transport miRNAs into vesicles that ultimately get transported as EVs outside the cell. The sorting of Ago2 into EVs was dependent on a signal initiated by KRAS through the MEK-ERK signaling pathway, a well-known pathway operating in normal cells in addition to cancerous ones. When the MEK-ERK signaling pathway was disrupted in cells with a cancerous version of KRAS, release of Ago2 into EVs was inhibited. The authors go on to identify specific phosphorylation sites responsible for inhibiting the association of Ago2 with cellular machinery and preventing secretion into EVs. Furthermore, disrupting these phosphorylation events also prevents the loading of specific miRNAs into EVs, showing that specific miRNAs, and not just random miRNAs, are selectively dependent on Ago2 for sorting into EVs. These results implicate Ago2 as a major player in a subset of RNAs sorting into EVs and identify a key regulatory signaling mechanism. Understanding and characterizing this pathway in further detail and in other systems will determine if these findings are conserved across cell types and systems. Ultimately, this may lead us to greater understanding of the normal function of extracellular RNA, as well as how these systems get disturbed or altered in diseases like cancer.
RNA carried in Extracellular Vehicles (EVs) and transported to neighboring or distant cells in the body can influence gene expression and other cellular process. However, not much is known about the mechanisms involved in the underlying processes. For example, how are EVs generated that carry these RNAs, and how are different types of RNAs, like microRNA (miRNA), sorted into different types of EVs? In this work, researchers shed light on some of the basic mechanisms and regulatory signals involved in sorting and transport of extracellular RNAs. Using well-characterized cancer cell lines, the researchers demonstrate that an RNA binding protein called Ago2, a component of the RNA-induced silencing complex (RISC), is responsible for helping transport miRNAs into vesicles that ultimately get transported as EVs outside the cell. The sorting of Ago2 into EVs was dependent on a signal initiated by KRAS through the MEK-ERK signaling pathway, a well-known pathway operating in normal cells in addition to cancerous ones. When the MEK-ERK signaling pathway was disrupted in cells with a cancerous version of KRAS, release of Ago2 into EVs was inhibited. The authors go on to identify specific phosphorylation sites responsible for inhibiting the association of Ago2 with cellular machinery and preventing secretion into EVs. Furthermore, disrupting these phosphorylation events also prevents the loading of specific miRNAs into EVs, showing that specific miRNAs, and not just random miRNAs, are selectively dependent on Ago2 for sorting into EVs. These results implicate Ago2 as a major player in a subset of RNAs sorting into EVs and identify a key regulatory signaling mechanism. Understanding and characterizing this pathway in further detail and in other systems will determine if these findings are conserved across cell types and systems. Ultimately, this may lead us to greater understanding of the normal function of extracellular RNA, as well as how these systems get disturbed or altered in diseases like cancer.
Reference:
KRAS-MEK Signaling Controls Ago2 Sorting into Exosomes. McKenzie, A.J., Hoshino, D., Hong, N.H., Cha, D.J., Franklin, J.L., Coffey, R.J., Patton, J.G. and Weaver, A.M., Cell Reports. 2016 Jan 24(1): 96-105.
Noninvasive Therapeutic Agents for Brain Cancers Show Promise in Mouse Studies
 Gaining access to the brain is a major obstacle for central nervous system drug development and delivery and non-invasive approaches are greatly needed. Research supported by the Extracellular RNA Communication Program is laying the foundation for the possibility of using extracellular RNAs (exRNAs) to treat brain cancers and other diseases. In this study, Dr. Zhang and colleagues use a mouse model to demonstrate the feasibility of treating brain-related diseases with intranasal delivery of RNA. They use a unique type of exosome-like nanoparticle, called a Grapefruit-derived Nanovector, to deliver an RNA of interest to the brain. These nanovectors can be generated from grapefruit plants in large quantities. Furthermore, Zhang et al. find that a specific RNA with potential therapeutic effects, miR17, can be effectively delivered to the brain using these grapefruit vectors without observable side effects in the mice. Remarkably, not only can the RNA be delivered to a target location effectively, it can also inhibit tumor growth in cell culture and in the mice. Although more work is needed to verify the mode of action and potential utility in humans, this study demonstrates the potential for an effective noninvasive therapeutic agent for the treatment of brain diseases.
Gaining access to the brain is a major obstacle for central nervous system drug development and delivery and non-invasive approaches are greatly needed. Research supported by the Extracellular RNA Communication Program is laying the foundation for the possibility of using extracellular RNAs (exRNAs) to treat brain cancers and other diseases. In this study, Dr. Zhang and colleagues use a mouse model to demonstrate the feasibility of treating brain-related diseases with intranasal delivery of RNA. They use a unique type of exosome-like nanoparticle, called a Grapefruit-derived Nanovector, to deliver an RNA of interest to the brain. These nanovectors can be generated from grapefruit plants in large quantities. Furthermore, Zhang et al. find that a specific RNA with potential therapeutic effects, miR17, can be effectively delivered to the brain using these grapefruit vectors without observable side effects in the mice. Remarkably, not only can the RNA be delivered to a target location effectively, it can also inhibit tumor growth in cell culture and in the mice. Although more work is needed to verify the mode of action and potential utility in humans, this study demonstrates the potential for an effective noninvasive therapeutic agent for the treatment of brain diseases.
Reference:
Grapefruit-derived Nanovectors Delivering Therapeutic miR17 Through an Intranasal Route Inhibit Brain Tumor Progression. Zhuang, X., Y. Teng, A. Samykutty, J. Mu, Z. Deng, L. Zhang, P. Cao, Y. Rong, J. Yan, D. Miller and H. G. Zhang Molecular Therapy. 2016 Jan 24(1): 96-105.
Image courtesy of the University of Louisville
Levels of microRNAs in the Blood can be Used to Monitor Development of Alcoholic Hepatitis
 Alcoholic hepatitis is a syndrome characterized by liver damage and inflammation resulting from chronic alcohol abuse. Researchers from the NIH Common Fund’s Extracellular RNA Communication program discovered increased levels of specific microRNAs in the blood of patients with alcoholic hepatitis, pointing to a new way to diagnose or monitor this disease. Dr. Gyongyi Szabo, current president of the American Association for the Study of Liver Diseases, and colleagues from the University of Massachusetts Medical School found that in both patients with alcoholic hepatitis and in mice fed a diet containing alcohol, there were increased levels of small vesicles known as exosomes in the bloodstream. One possible cargo of exosomes are microRNAs, and the researchers determined that several microRNAs were increased in the blood from alcohol-fed mice and hepatitis patients. Dr. Szabo and colleagues also demonstrated the blood levels of one specific microRNA, miR-192, can be used to accurately distinguish patients with alcoholic hepatitis from healthy controls. This research has the potential to improve the ability of doctors to diagnose this important health condition and to monitor a patient’s response to treatment.
Alcoholic hepatitis is a syndrome characterized by liver damage and inflammation resulting from chronic alcohol abuse. Researchers from the NIH Common Fund’s Extracellular RNA Communication program discovered increased levels of specific microRNAs in the blood of patients with alcoholic hepatitis, pointing to a new way to diagnose or monitor this disease. Dr. Gyongyi Szabo, current president of the American Association for the Study of Liver Diseases, and colleagues from the University of Massachusetts Medical School found that in both patients with alcoholic hepatitis and in mice fed a diet containing alcohol, there were increased levels of small vesicles known as exosomes in the bloodstream. One possible cargo of exosomes are microRNAs, and the researchers determined that several microRNAs were increased in the blood from alcohol-fed mice and hepatitis patients. Dr. Szabo and colleagues also demonstrated the blood levels of one specific microRNA, miR-192, can be used to accurately distinguish patients with alcoholic hepatitis from healthy controls. This research has the potential to improve the ability of doctors to diagnose this important health condition and to monitor a patient’s response to treatment.
Reference:
Increased number of circulating exosomes and their microRNA cargos are potential novel biomarkers in alcoholic hepatitis. Momen-Herav Fi, Saha B, Kodys K, Catalano D, Satishchandran A and Szabo G. J Transl Med. 2015 August 12. 13:261.
Glioblastoma Cells use Extracellular RNA to Influence Normal Cells in the Brain
 In research supported by the Common Fund’s Extracellular RNA Communication program, Dr. Xandra Breakefield and colleagues have discovered that glioblastoma tumor cells can manipulate healthy cells around them through the release of extracellular RNA (exRNAs). Glioblastoma is the most common form of brain cancer and even with standard treatments is associated with very poor prognoses. Glioblastoma tumor cells manipulate healthy brain cells to become more supportive of tumor growth, but it is unclear how this occurs. This study found that one way glioblastoma cells manipulate healthy cells is by the release of small vesicles containing exRNAs that are in turn taken up by healthy cells. Among the vesicle bound exRNAs are several highly abundant microRNAs, which can alter the rate of protein production and thus change growth promoting properties of the healthy cells that take them up. Using fluorescently tagged cells and vesicles, Dr. Breakefield and colleagues directly visualized transfer of these vesicles from tumor to healthy cells in the brains of mice with glioblastomas. In addition, by isolating the target cells, the researchers demonstrated changes in protein composition, consistent with the hypothesis that the transferred microRNAs caused a functional change in the target cells. These studies suggest that exRNA may play an instructive role in promoting an environment conducive to growth of a glioblastoma, and therapeutics targeting specific exRNAs may warrant further exploration in developing treatments for glioblastoma.
In research supported by the Common Fund’s Extracellular RNA Communication program, Dr. Xandra Breakefield and colleagues have discovered that glioblastoma tumor cells can manipulate healthy cells around them through the release of extracellular RNA (exRNAs). Glioblastoma is the most common form of brain cancer and even with standard treatments is associated with very poor prognoses. Glioblastoma tumor cells manipulate healthy brain cells to become more supportive of tumor growth, but it is unclear how this occurs. This study found that one way glioblastoma cells manipulate healthy cells is by the release of small vesicles containing exRNAs that are in turn taken up by healthy cells. Among the vesicle bound exRNAs are several highly abundant microRNAs, which can alter the rate of protein production and thus change growth promoting properties of the healthy cells that take them up. Using fluorescently tagged cells and vesicles, Dr. Breakefield and colleagues directly visualized transfer of these vesicles from tumor to healthy cells in the brains of mice with glioblastomas. In addition, by isolating the target cells, the researchers demonstrated changes in protein composition, consistent with the hypothesis that the transferred microRNAs caused a functional change in the target cells. These studies suggest that exRNA may play an instructive role in promoting an environment conducive to growth of a glioblastoma, and therapeutics targeting specific exRNAs may warrant further exploration in developing treatments for glioblastoma.
Reference:
Directly visualized glioblastoma-derived extracellular vesicles transfer RNA to microglia/macrophages in the brain. van der Vos KE, Abels ER, Zhang X, Lai C, Carrizosa E, Oakley D, Prabhakar S, Mardini O, Crommentuijn MH, Skog J, Krichevsky AM, Stemmer-Rachamimov A, Mempel TR, El Khoury J, Hickman SE, Breakefield XO. Neuro Oncol. 2015 Oct 3.
NIH Consortium is Advancing our Understanding of Extracellular RNA Communication
 The Common Fund-supported Extracellular RNA Communication Consortium (ERCC) has published six manuscripts in a recent special issue of the Journal of Extracellular Vesicles, providing the scientific community with information about the expected outcomes of this new scientific program and detailing major progress to date. The overarching goal of the ERCC is to generate a comprehensive community resource to share fundamental scientific discoveries, protocols, and innovative tools and technologies related to extracellular (“outside the cell”) RNA. An emerging field of research, extracellular RNA is now known to play a role in newly discovered mechanisms of cell-to-cell communication. In recognition of the paradigm-shifting potential of extracellular RNA communication in human health and disease, the NIH is supporting a comprehensive program to catalyze research in this area. The Extracellular RNA Communication program spans basic and translational research, building a foundation of fundamental biological principles of exRNA communication and understanding of exRNAs in healthy people, and then leveraging this new knowledge to develop biomarkers and therapies for a wide variety of diseases and conditions.
The Common Fund-supported Extracellular RNA Communication Consortium (ERCC) has published six manuscripts in a recent special issue of the Journal of Extracellular Vesicles, providing the scientific community with information about the expected outcomes of this new scientific program and detailing major progress to date. The overarching goal of the ERCC is to generate a comprehensive community resource to share fundamental scientific discoveries, protocols, and innovative tools and technologies related to extracellular (“outside the cell”) RNA. An emerging field of research, extracellular RNA is now known to play a role in newly discovered mechanisms of cell-to-cell communication. In recognition of the paradigm-shifting potential of extracellular RNA communication in human health and disease, the NIH is supporting a comprehensive program to catalyze research in this area. The Extracellular RNA Communication program spans basic and translational research, building a foundation of fundamental biological principles of exRNA communication and understanding of exRNAs in healthy people, and then leveraging this new knowledge to develop biomarkers and therapies for a wide variety of diseases and conditions.
In this special issue, the ERCC describes how its data, resources, and protocols can be used by the wider scientific community, with the goal of advancing broad participation in exRNA research. They are developing a community resource, the exRNA Portal which provides access to exRNA resources and products developed by the ERCC.
Also within this special issue, the current landscape of exRNA research is described, and the progress to date within the ERCC is detailed. In a manuscript addressing biogenesis, delivery, and function of extracellular RNA, the authors highlight projects investigating the role of exRNAs in glioblastoma and colorectal cancer, exploring fundamental processes of exRNA generation and release, and developing genetic and cellular models to investigate exRNA communication. A manuscript about the Consortium’s biomarker efforts describes examining diverse body fluids such as plasma, serum, urine, and cerebrospinal fluid to detect markers for diseases as varied as cancer, cardiovascular disease, kidney disease, neurodegenerative disease, hemorrhage, and placental disorders. Addressing the large diversity and volume of exRNA data that will be included in the exRNA Portal, another manuscript outlines the Consortium’s strategy for data integration using metadata, biomedical ontologies, and linked data technologies. Another manuscript explores our evolving understanding of the functional effects of extracellular vesicles, including their role in digestive, immune, and cardiovascular function; recovery from injury or disease; cancer progression; and use as engineered therapeutics for brain diseases. Finally, one manuscript identifies the need for robust and standardized methods for collecting and processing biofluids, separation of different types of exRNA-containing particles, and isolation and analysis of exRNAs, as well as the next steps for identifying high-priority methodological needs that would promote rapid advancements and consistency in the exRNA field.
Collectively, these papers outline a clear and comprehensive vision for the ERCC, while providing the broader scientific community with information about important exRNA resources they can use to support their own research.
References:
The NIH Extracellular RNA Communication Consortium. Ainsztein et al. Journal of Extracellular Vesicles. 2015.
Biogenesis, delivery, and function of extracellular RNA. Patton et al. Journal of Extracellular Vesicles. 2015.
Potential functional applications of extracellular vesicles: a report by the NIH Common Fund Extracellular RNA Communication Consortium. Quesenberry et al. Journal of Extracellular Vesicles. 2015.
Extracellular RNAs: development as biomarkers of human disease. Quinn et al. Journal of Extracellular Vesicles. 2015.
Integration of extracellular RNA profiling data using metadata, biomedical ontologies, and Linked Data technologies. Subramanian et al. Journal of Extracellular Vesicles. 2015.
Meeting report: Discussion and preliminary findings on Extracellular RNA measurement methods of laboratories in the NIH Extracellular RNA Communication Consortium. Laurent et al. Journal of Extracellular Vesicles. 2015.
Read the blog post about these papers on the exRNA Portal
Insights into Potential exRNA Biomarkers for Breast Cancer
Researchers supported by the Common Fund’s Extracellular RNA Communication program are gaining new insight into the potential for some types of extracellular RNA called microRNA (miRNA) to influence cancer progression. The research suggests that cancer cell exosomes, vesicles that are secreted by cells and present in many biological fluids, and the associated mature miRNA they carry are involved in tumor formation. Since only cancer exosomes contain certain proteins, these could serve as important diagnostic markers that may be more useful in detection of cancers compared to some current diagnostic approaches such as imaging. This important distinction may also give insight into the design of novel cancer therapeutics. Though the functional roles of miRNAs remain largely unknown, researchers are beginning to understand their importance in influencing transcription and tumor progression in target cells. In this study, researchers found that breast cancer cells secrete exosomes packed with the necessary proteins to process miRNAs into their mature form, while exosomes from normal non-cancerous cells lacked this ability. The scientists also showed that when cancer exosomes were combined with normal human breast cells and injected into the mammary tissue of mice, the injected cells became cancerous and formed tumors. Conversely, exosomes from sera of healthy donors had no tumorigenic effect. This research suggests a new cell-independent mechanism by which secreted exosomes from cancerous cells are able to process miRNA that can influence and signal non-cancerous cells to become cancerous, demonstrating that RNA released from one part of the body may have the potential to influence cells in distant parts of the body.
are gaining new insight into the potential for some types of extracellular RNA called microRNA (miRNA) to influence cancer progression. The research suggests that cancer cell exosomes, vesicles that are secreted by cells and present in many biological fluids, and the associated mature miRNA they carry are involved in tumor formation. Since only cancer exosomes contain certain proteins, these could serve as important diagnostic markers that may be more useful in detection of cancers compared to some current diagnostic approaches such as imaging. This important distinction may also give insight into the design of novel cancer therapeutics. Though the functional roles of miRNAs remain largely unknown, researchers are beginning to understand their importance in influencing transcription and tumor progression in target cells. In this study, researchers found that breast cancer cells secrete exosomes packed with the necessary proteins to process miRNAs into their mature form, while exosomes from normal non-cancerous cells lacked this ability. The scientists also showed that when cancer exosomes were combined with normal human breast cells and injected into the mammary tissue of mice, the injected cells became cancerous and formed tumors. Conversely, exosomes from sera of healthy donors had no tumorigenic effect. This research suggests a new cell-independent mechanism by which secreted exosomes from cancerous cells are able to process miRNA that can influence and signal non-cancerous cells to become cancerous, demonstrating that RNA released from one part of the body may have the potential to influence cells in distant parts of the body.
Read the article in Cancer Cell
Reference:
Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Melo SA, Sugimoto H, O'Connell JT, Kato N, Villanueva A, Vidal A, Qiu L, Vitkin E, Perelman LT, Melo CA, Lucci A, Ivan C, Calin GA, Kalluri R. Cancer Cell. 2014 Nov 12. 26(5): 707-721.
Extracellular RNA Communication Researcher Discovers "Treasure in Saliva"
Research supported by the Common Fund’s Extracellular RNA Communication program is laying the foundation for using extracellular RNAs (exRNAs) in saliva to diagnose a variety of diseases, such as cancer, diabetes, autoimmune disorders, and potentially many more. Dr. David Wong and colleagues have conducted the most comprehensive analysis of human exRNAs in saliva to date, and have made several interesting discoveries. One surprising finding is the discovery of approximately 400 circular RNAs in saliva. Circular RNAs were only recently discovered to exist within cells and tissues, and this study marks the first discovery of circular RNAs in any body fluid. The researchers also discovered that saliva contains a significant number of piwi-interacting RNAs (piRNAs), whose functions are largely unknown. In contrast, blood and other body fluids contain very few piRNAs, suggesting that the salivary piRNAs did not originate from RNAs in the blood, and may have originated instead from skin or stem cells within the oral cavity. MicroRNAs (miRNAs), which play important roles in gene regulation, were also found in saliva, and show similar variability between individuals in the saliva samples compared to miRNAs from blood samples or within cells. This result suggests salivary miRNAs may have potential as stable biomarkers that can distinguish between individuals, with the advantage of being easily accessible compared to other body fluids. Together, this research is the first step for future studies on the biological functions of exRNAs in saliva, and opens the door to using salivary exRNAs as non-invasive biomarkers for a number of diseases.
foundation for using extracellular RNAs (exRNAs) in saliva to diagnose a variety of diseases, such as cancer, diabetes, autoimmune disorders, and potentially many more. Dr. David Wong and colleagues have conducted the most comprehensive analysis of human exRNAs in saliva to date, and have made several interesting discoveries. One surprising finding is the discovery of approximately 400 circular RNAs in saliva. Circular RNAs were only recently discovered to exist within cells and tissues, and this study marks the first discovery of circular RNAs in any body fluid. The researchers also discovered that saliva contains a significant number of piwi-interacting RNAs (piRNAs), whose functions are largely unknown. In contrast, blood and other body fluids contain very few piRNAs, suggesting that the salivary piRNAs did not originate from RNAs in the blood, and may have originated instead from skin or stem cells within the oral cavity. MicroRNAs (miRNAs), which play important roles in gene regulation, were also found in saliva, and show similar variability between individuals in the saliva samples compared to miRNAs from blood samples or within cells. This result suggests salivary miRNAs may have potential as stable biomarkers that can distinguish between individuals, with the advantage of being easily accessible compared to other body fluids. Together, this research is the first step for future studies on the biological functions of exRNAs in saliva, and opens the door to using salivary exRNAs as non-invasive biomarkers for a number of diseases.
Read the news release from the University of California Los Angeles
Read the article in Science Daily
Reference:
The Landscape of MicroRNA, Piwi-Interacting RNA, and Circular RNA in Human Saliva. Bahn JH, Zhang Q, Li F, Chan T-M, Lin X, Kim Y, Wong D, and Xiao X. Clinical Chemistry, Jan 2015, 61 (1) 221-230.
Extracellular RNA Shows Promise in Treating Multiple Sclerosis and Other Neurological Diseases
Researchers in the Common Fund’s Extracellular RNA Communication program  have discovered a potential treatment for multiple sclerosis (MS), a devastating neurological disorder characterized by muscle weakness, vision problems, difficulty with balance and coordination, and sometimes paralysis. Dr. Richard Kraig and colleagues from the University of Chicago are investigating the therapeutic potential of exosomes, small particles containing biologically active molecules such as RNA and proteins, which are released from cells to travel throughout the body and affect other cells at a distance. Dr. Kraig’s research shows immune cells can be stimulated to produce exosomes that promote formation of myelin to restore the protective insulation around nerve fibers that is damaged in MS. These exosomes contain small pieces of genetic material called microRNAs. Some microRNAs in the exosomes influence immature brain cells to develop into myelin-making cells called oligodendrocytes. Other microRNAs protect against inflammation, thought to contribute to myelin damage in MS. Treatment with exosomes containing these microRNAs increases myelin in both healthy rodent brains and in rat models of demyelination that mimic MS. Importantly, a nasal spray containing exosomes with microRNAs was found to increase myelin in rat brains, suggesting that this type of treatment may be easily administered. In related research, Dr. Kraig and colleagues found that microRNAs in exosomes from young animals and animals living in environmentally enriched conditions also promote myelination, suggesting multiple factors may influence production of microRNA-containing exosomes with therapeutic potential. Further studies will be needed to determine whether exosomal microRNAs can be used to treat patients with MS, but these early studies are a promising first step in developing microRNA-based therapeutics for MS and possibly many other neurological diseases and conditions.
have discovered a potential treatment for multiple sclerosis (MS), a devastating neurological disorder characterized by muscle weakness, vision problems, difficulty with balance and coordination, and sometimes paralysis. Dr. Richard Kraig and colleagues from the University of Chicago are investigating the therapeutic potential of exosomes, small particles containing biologically active molecules such as RNA and proteins, which are released from cells to travel throughout the body and affect other cells at a distance. Dr. Kraig’s research shows immune cells can be stimulated to produce exosomes that promote formation of myelin to restore the protective insulation around nerve fibers that is damaged in MS. These exosomes contain small pieces of genetic material called microRNAs. Some microRNAs in the exosomes influence immature brain cells to develop into myelin-making cells called oligodendrocytes. Other microRNAs protect against inflammation, thought to contribute to myelin damage in MS. Treatment with exosomes containing these microRNAs increases myelin in both healthy rodent brains and in rat models of demyelination that mimic MS. Importantly, a nasal spray containing exosomes with microRNAs was found to increase myelin in rat brains, suggesting that this type of treatment may be easily administered. In related research, Dr. Kraig and colleagues found that microRNAs in exosomes from young animals and animals living in environmentally enriched conditions also promote myelination, suggesting multiple factors may influence production of microRNA-containing exosomes with therapeutic potential. Further studies will be needed to determine whether exosomal microRNAs can be used to treat patients with MS, but these early studies are a promising first step in developing microRNA-based therapeutics for MS and possibly many other neurological diseases and conditions.
References:
IFNγ-stimulated dendritic cell exosomes as a potential therapeutic for remyelination. Pusic AD, Pusic KM, Clayton BLL, and Kraig RP. Journal of Immunology, Jan. 15, 2014; 266(1-2): 12-23. PMID: 24275061.
Youth and environmental enrichment generate serum exosomes containing miR-219 that promote CNS myelination. Pusic AD and Kraig RP. Glia, Feb. 2014; 62(2): 284-299. PMID: 24339157.
Read about this story in the news:
Naturally Occurring Packets Show Promise for Protecting Nerve Fibers in the Brain
Remyelination: Are Exosomes Containing microRNA the Answer?
NIH Common Fund Issues First Awards in Extracellular RNA Communication!
The NIH is supporting 24 collaborative, multidisciplinary awards to explore a novel cell communication process.
Read the press release
View the Funded Research


